Data Provenance¶
One of the goals of PyARPES is to allow for reproducible and easily introspectable scientific analysis. It should be easy for an experimenter to go back to prior analysis notebooks or to figure and understand how it was made. Similarly, reproducible science requires that others can do the same.
Even hygienic use of Jupyter notebooks is insufficient to achieve true reproducible analysis because
Code can change
Data is mutable and a precise record of changes to data is not recorded
Finished analysis products like figures are not linked to analysis
PyARPES attempts to address these problems by maintaining a record of actions performed on data including functions, software version, and parameters, when you use the builtin analysis routines.
PyARPES also configures Jupyter and IPython to log all of your Jupyter
sessions to logs/{notebook-name}_{date}_{time}.log. You can opt out
of this by using the LOGGING_ENABLED feature switch in
arpes.config.CONFIG.
This record is used to produce a lineage for the data involved in any
figure created using the PyARPES standard figures, or using the PyARPES
savefig wrapper. Additionally, each element in this data provenance
record stores the most recent cells run inside an IPython or Jupyter
session, if available, in order to make available the outside-of-library
context for this item in the provenance.
PyARPES also provides tools that make it simple to record and maintain data provenance for your analysis code.
Format for data provenance¶
You can use .S.history to obtain the data provenance for data you
work with in PyARPES. Provenance is available on xr.DataArray and
xr.Dataset instances and takes a nested list format, with IDs used
for references.
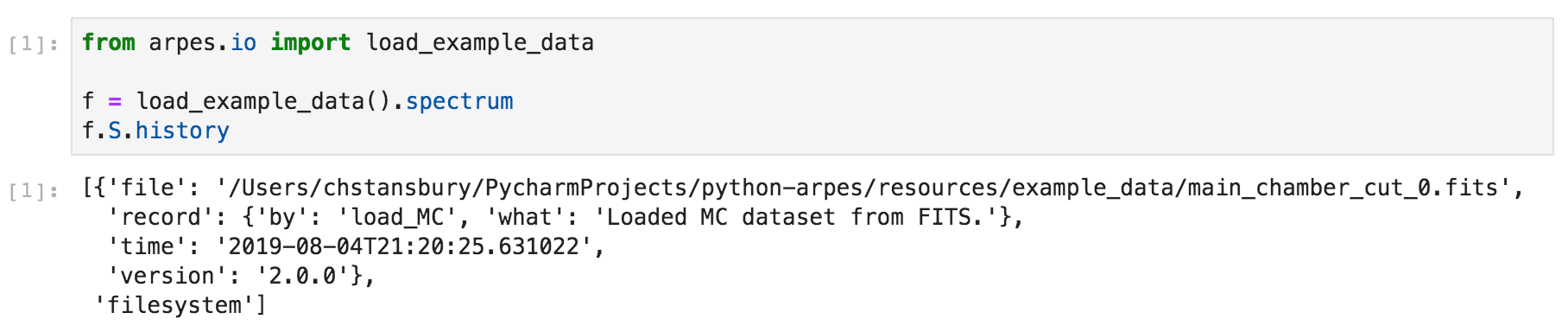
Example data provenance in PyARPES¶
Most analysis routines that consume and produce xarray based data
will update this provenance record.
Here we can see that converting to momentum space includes a new item indicating the conversion.
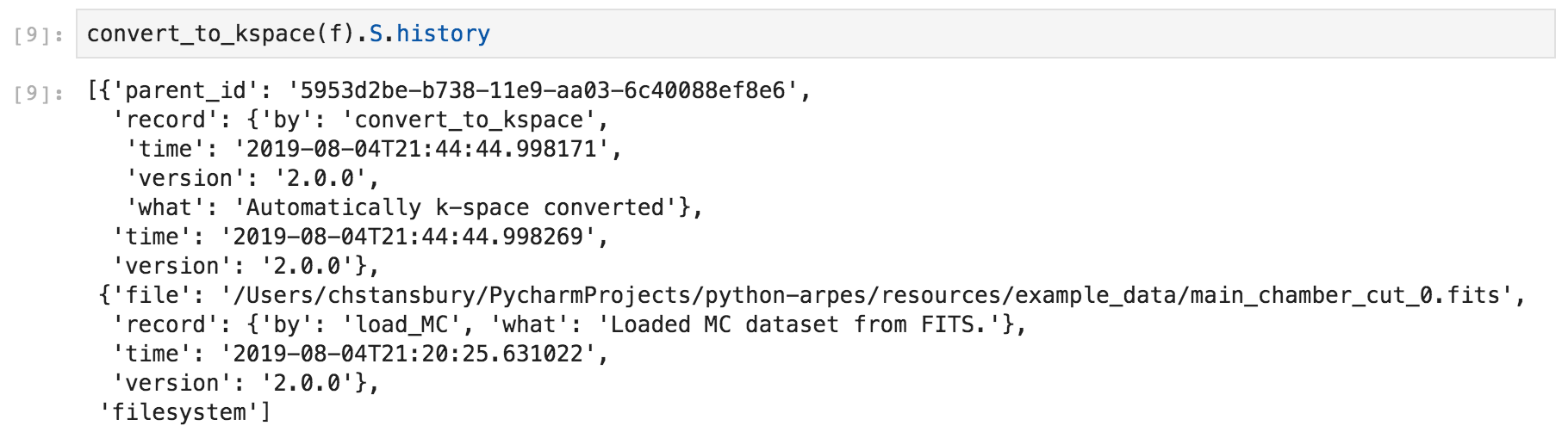
Example data provenance in PyARPES¶
Outputting provenance when plotting¶
After you’ve performed some analysis, you will want to produce plots, and you will want the provenance to stick around with the figure.
The built-in plotting scripts in PyARPES will save both the output figure, and a record of the data provenance.
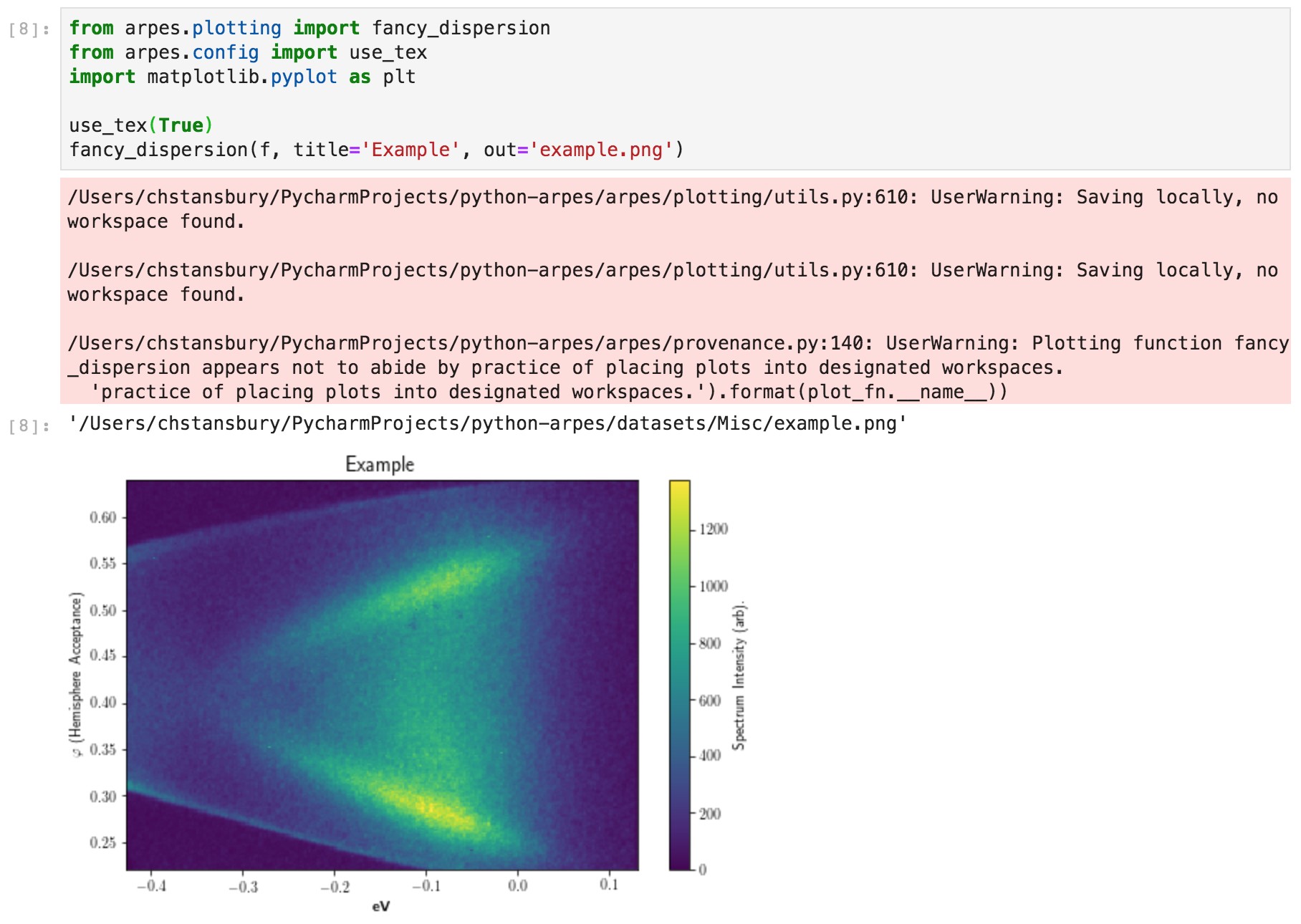
Example data provenance in PyARPES¶
As an example, here we plot an ARPES cut: the figure is saved as “example.png”, while the provenance for the data in the figure is recorded in the file “example.png.provenance.json”.
After plotting, the provenance file contains:
{
"VERSION": "2.0.0",
"args": [
{
"version": "2.0.0",
"record": {
"what": "Loaded MC dataset from FITS.",
"by": "load_MC"
},
"time": "2019-08-04T21:20:25.631022",
"parents_provenance": "filesystem",
"file": "/Users/chstansbury/PycharmProjects/python-arpes/resources/example_data/main_chamber_cut_0.fits"
}
],
"kwargs": {},
"time": "2019-08-04T21:43:39.581877",
"name": "fancy_dispersion"
}
Custom plots¶
Often you need to customize or write a one off plot. If instead of using
Matplotlib’s savefig you use arpes.plotting.savefig, then the
data= parameter allows you to request a provenance record for data
in your custom plot.
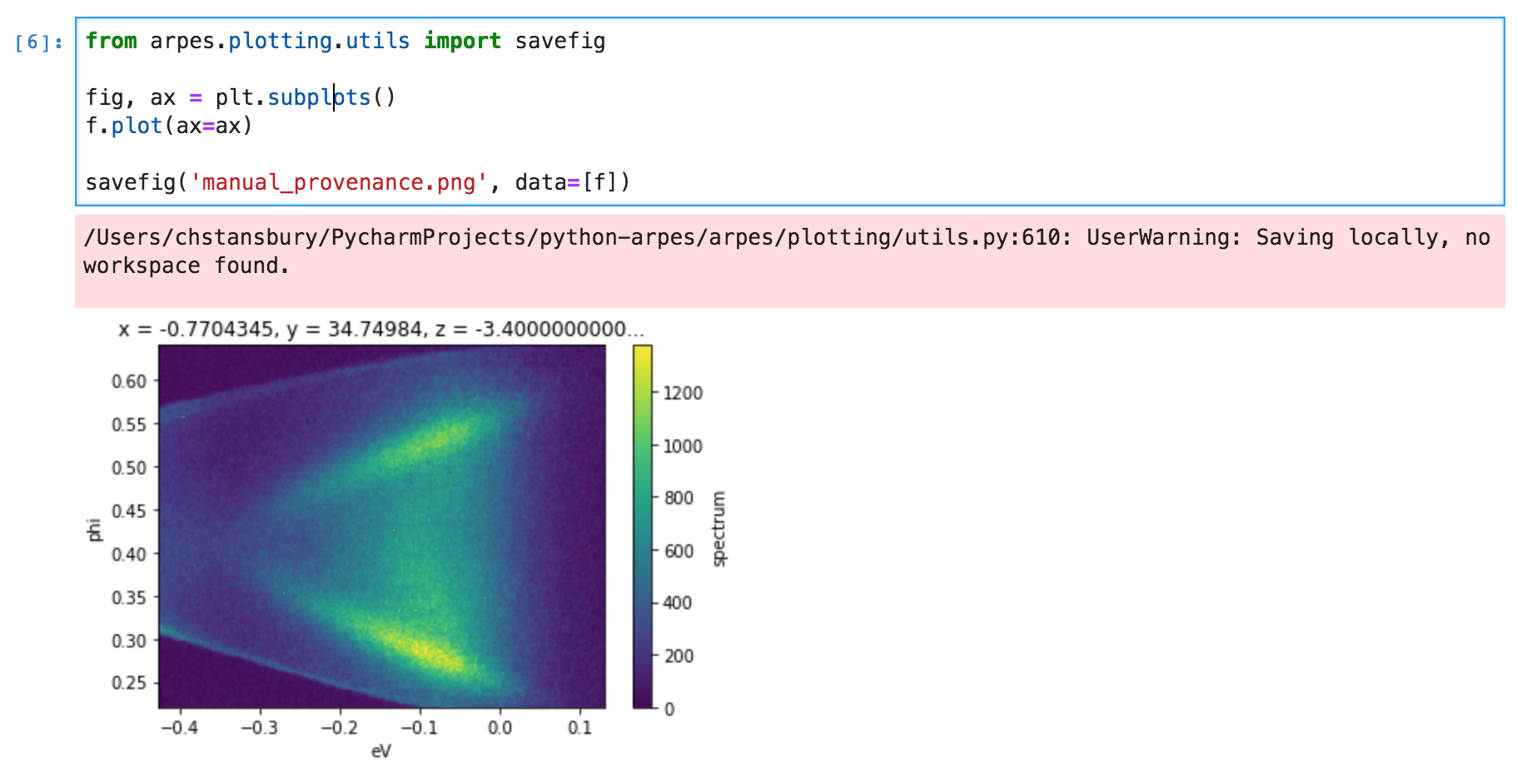
Example data provenance in PyARPES¶
This records a provenance more or less identical to the previous example.
Using provenance in your code¶
PyARPES makes it very straightforward to opt into recording provenance.
If you write an analysis function, the simplest option is to use the
update_provenance decorator, which introspects data passed to and
returned from your function at runtime and updates provenance
appropriately.
Additionally, because PyARPES includes recent Juypyter cell evaluations in the record, you shouldn’t find the need to be very defensive in your use of manual provenance tracking, although you should wrap code that forms part of your more permanent analysis repetoire.
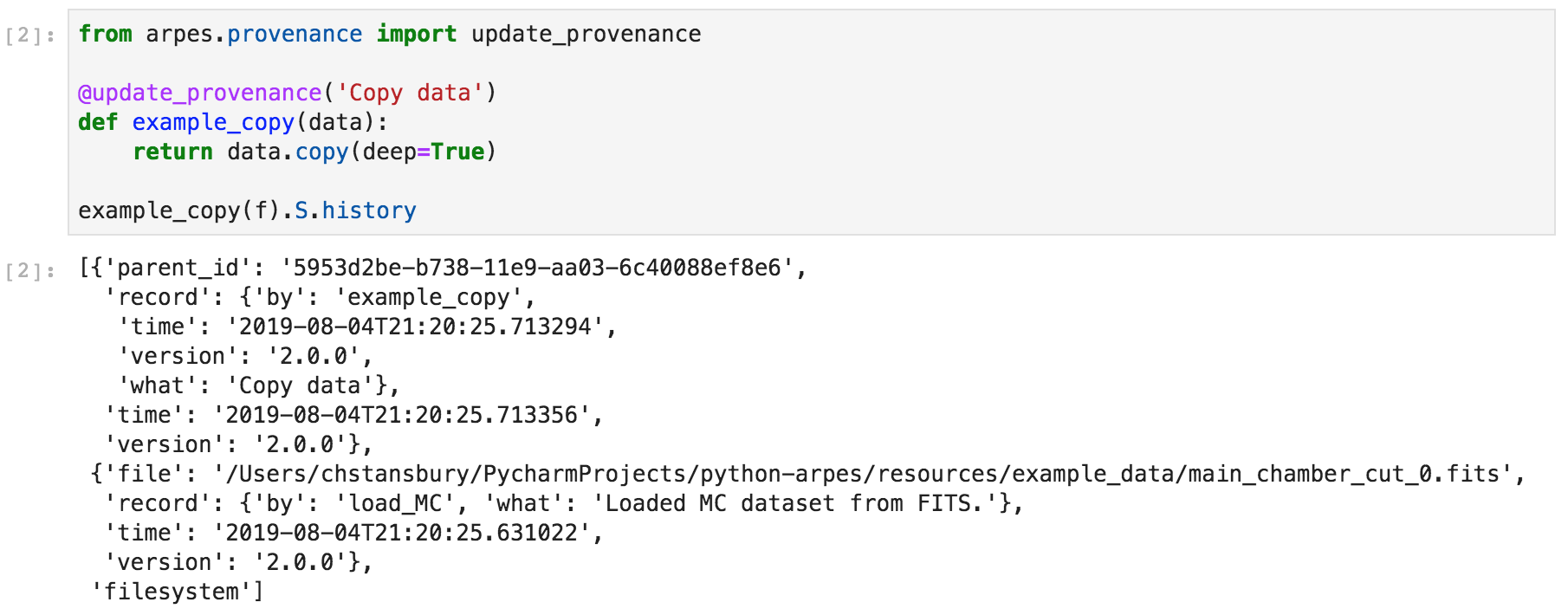
Example data provenance in PyARPES¶
If you need more control, you can also manually modify the provenance record
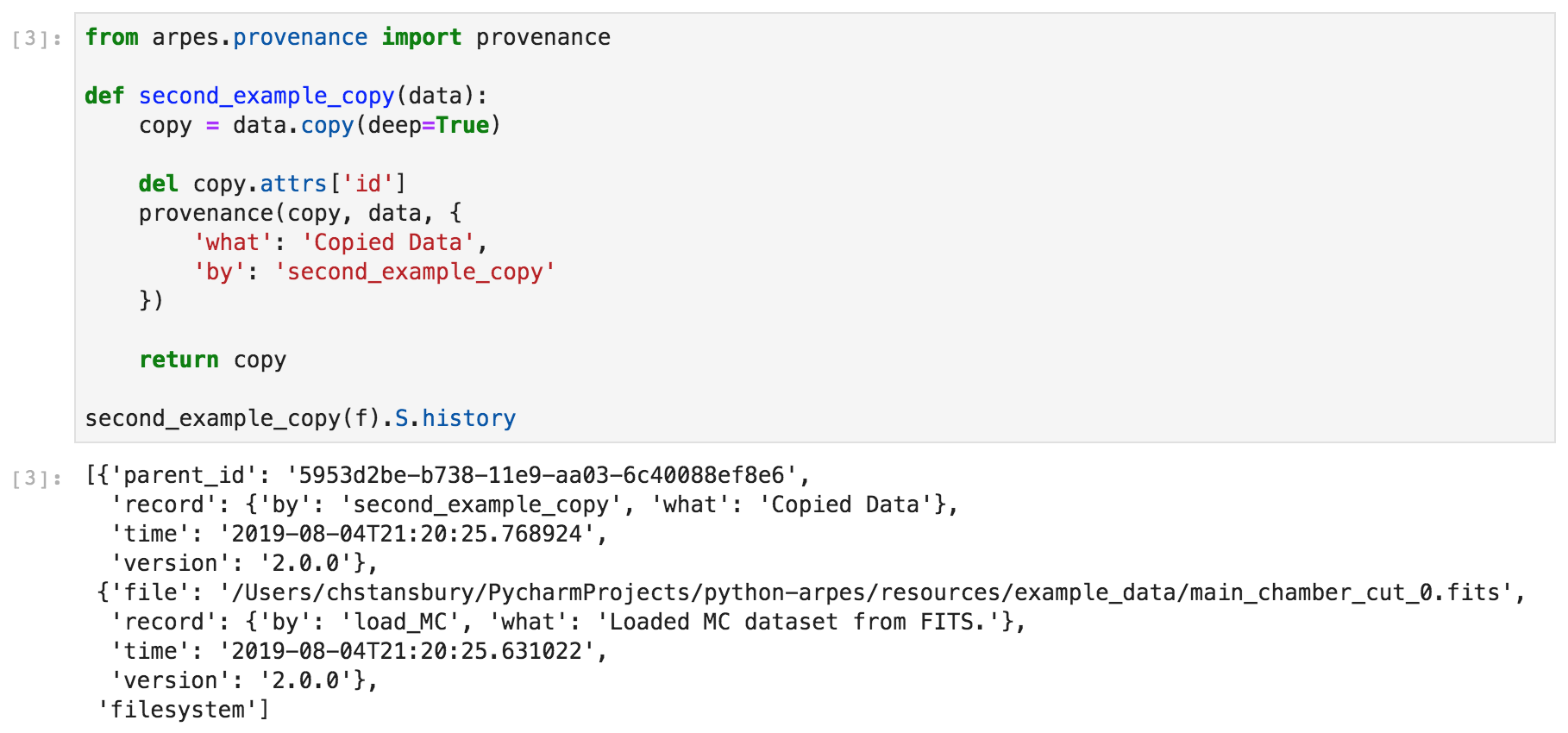
Example data provenance in PyARPES¶